Dr Vassili Kovalev, at the United Institute of Informatics Problems of the National Academy of Sciences of Belarus, knows all about the importance of collaboration. As head of the Department of Biomedical Image Analysis, he and his team found that only by looking outwards could they overcome an obstacle in their research.
Dr Kovalev and his team specialise in the use of Biomedical Image Analysis and Deep Learning technologies in medical diagnosis and treatment. But the use of large amounts of personal medical data involved in the training of Deep Neural Networks (DNNs) had presented the scientific community with a number of ethical, legal, and security challenges.
They explored two avenues to overcome this: the development of suitable artificial image data that was indistinguishable from real samples while training the Deep Learning models; and raising security levels to confront so-called ‘adversarial attacks’ to DNNs.
So, the team set about applying Deep Learning methods to solve conventional computerised disease diagnosis tasks. At the same time, they started developing new methods and software for the automatic generation of artificial biomedical images and studying methods of crafting malicious images used in adversarial attacks and suitable ways of defense.
Generating medical images of people who don’t exist
Lately, there has been enormous progress in new methods of image analysis and classification, which is based on Deep Learning approaches. Such methods often provide very good results. But the substantial drawback is that they use very large collections of suitably labelled biomedical images for training neural networks.”
These large collections, even when completely depersonalised, are still questionable from an ethical and a security standpoint. But Kovalev and his team developed a solution. They came up with a method using Generative Adversarial Networks to automatically generate medical images, which could be used as a substitute for real ones.
Once the high visual quality and adequate quantitative properties of the artificial images were achieved, they could be used for both the training of Deep Learning models, but also for educational and illustrative purposes to assure software quality. “Using this case,” continues Dr Kovalev, “scientists would be free to do any kind of research based on such artificial data.”
2 million
chest X-ray images
from private databases, used for training neural networks.
10000
patients' 3D computed tomography images
from private databases, used for training neural networks.
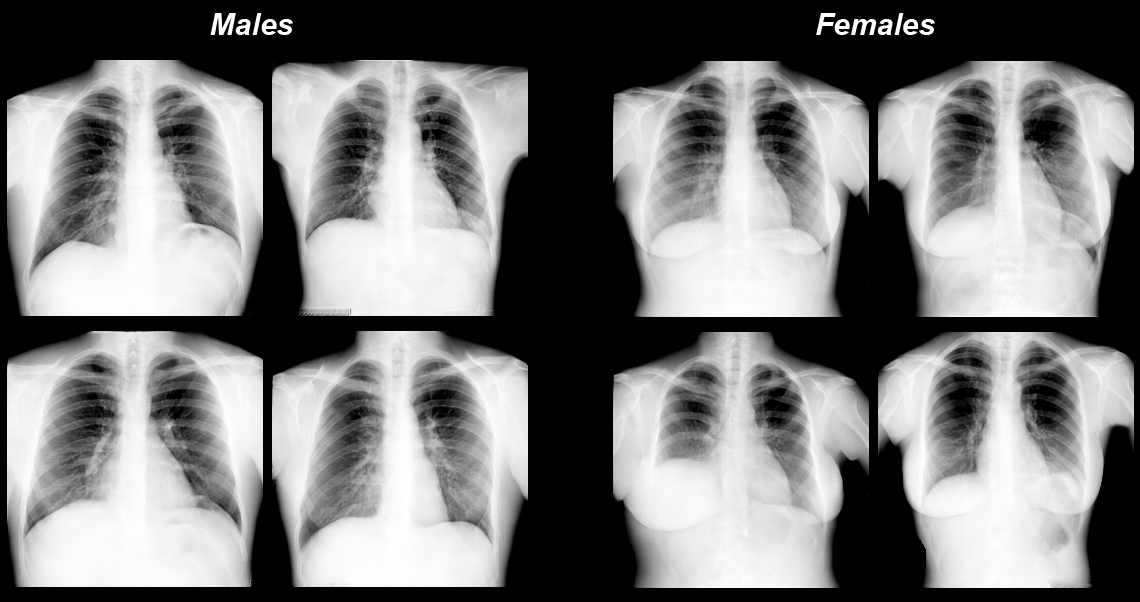
So far so good. But the image generation research was based on very large private medical image databases with restricted access. These included a chest X-Ray database containing about 2,000,000 natively digital images, a database of 3D computed tomography images of 10,000 patients, and a database of histology images used for cancer diagnoses. Some slides were several Terabytes in size.
All this needed high computational power, something the Belarussian team were severely lacking. So, they started looking into their other area of research – the problem of security surrounding biomedical image analysis technologies in the real disease diagnosis set-up.
Battling ‘adversarial attacks’
So, what was the problem? Researchers discovered a security phenomenon – if a tiny modification was made to an original image, someone could force well-trained DNN to make a wrong decision. In the context of the medical diagnosis processes, this was a very dangerous situation. A healthy person with a slightly amended image could receive results that were considered suspicious for cancer.
It was experimentally proven that virtually any deep neural network could be forced to make the wrong decision.”
So, the Belarussian researchers conducted a number of computational Deep Learning experiments to study the vulnerability of different medical images in order to protect computerised disease diagnosis systems from potential attacks by malicious medical images.
Again, they encountered the same blocker – the lack of high computational power. What they needed was access to HPC (high performance computing).
So how did they gain access to HPC?
In both cases, to operate with such a big amount of data and to use state-of-the-art image analysis methods and software the team needed very powerful computational resources.
“We realised quickly it would be very important to join forces with other research facilities,” admits Dr Kovalev. The scientists applied to the Enlighten Your Research (EYR) programme, an initiative run by the EU-funded Eastern Partnership Connect (EaPConnect) project . This project aims to bring together researchers, academics and students across Armenia, Azerbaijan, Belarus, Georgia, Moldova Ukraine, so they can collaborate with each other and their peers around the world by connecting via highly reliable high-capacity networks and dedicated online services.
Kovalev’s team qualified for EYR and the results were immediate. The computational bottleneck was removed. They gained access to European HPC resources through the infrastructure provided by the Belarusian national R&E network BASNET and the pan-European R&E network GÉANT.
Finally, they could proceed with their work. They can now perform the majority of their experiments and share the new knowledge and generated artificial images with European partners.